|
A simple example is shown in Figure 2 where a 1 kHz signal with the amplitude of 10 mV is shown both in the Scope tool and streamed via Plotter. Continuous StreamingĬontinuous Scope streaming allows you to stream any of the two scope channels directly on the Plotter with the adjustable streaming rate of 1.83 kHz up to 3.75 MHz. In the Frequency domain, on the other hand, you get an increased frequency resolution (FR) that you simply get by dividing the chosen Sample Rate with the given sample length, FR = SR/L, which for the above example amounts to 91 mHz. Red framed box on the right depicts the Node Tree selection available for the MFLI with MF-DIG option.įor a given sample length (L) and Sample Rate (SR) the scope shot length of time (LT) can be calculated as: LT = L/SR, so for a SR= 234 Sa/s and the maximum L=2,56 MSa we get LT= 10,94 s. By adding more options, such as the MF-MD option, the Node tree is growing accordingly.įigure 1: A screen shot of the Scope time domain trace showing simultaneously the Voltage (blue) and the Current (red) Input signals. The Input Node Tree for an MFLI unit with MF-DIG option is shown in the right red box of Figure 1. However you also get an access to the Demodulator data which you can depict as a signal input for the Scope channel. The Scope can now serve as a two channel time/frequency domain tool where you can monitor current (I) and voltage (V) inputs at the same time, and also any other signal port such as Aux (In/Out)s, Trigger (In/Out)s, etc. With the MF-DIG option installed several advanced measurement opportunities open up. 
With a basic MFLI unit, the Scope comes with the possibility to monitor one input channel at at time with a variable sampling rate from 60 MSa/s down to 916 Sa/s and maximum length of 16 k points. Let's see how this is done by looking at what MFLI Scope with MF-DIG option offers for measurements in the time and frequency domain. This has proved crucial for performing the cross-correlation measurements to lower the noise floor using the two MFLIs or a single UHFLI (see this blog post too). When we enable the MF-DIG Digitizer option on an MFLI we get a chance to perform a higher resolution noise spectra and connect the lock-in measurement with that of the scope by advanced triggering capabilities. In this blog, we see how this is possible and cover several important advantages of having the Digitizer Option that makes the Scope an invaluable part of your measurements.Īs all Zurich Instruments' lock-ins come with an oscilloscope that utilizes the digitized signal at the input, the raw signal in the time and frequency domain is always available. When we make new devices, explore new materials, and research a new topic we get a chance to learn from the noise figures and use the scope as an integral part of the research. Throughout the measurement, detailed noise spectra reveal critical changes in the physics of the experiment but an external oscilloscope is likely to interfere with the measurements. When the setup is well characterized the measurements are carried out at a single frequency and measured against external parameters. 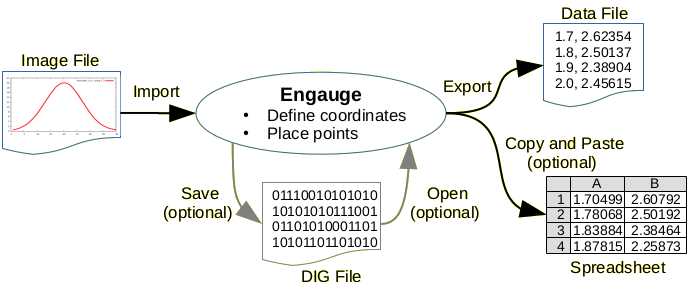
Lock-in amplifiers are at their best when rejecting the noise and increasing the signal-to-noise ratio. Non-Contact Atomic Force Microscopy (NC-AFM) Multi-Frequency Atomic Force Microscopy (MF-AFM) Tunable Diode Laser Absorption Spectroscopy Magnetometry with Ensembles of NV Centers Either a completed version of my function or any other method that achieves my goal would be extremely helpful.Quantum Computing with Superconducting Qubits In theory, this function should do what I want to achieve, but when tested it doesn't yield the expected results. Hd_pixel = np.ravel(hd_pixel) #turns 2D array into 1D to be able to compute average 
Grid_shape: an array of length 2 which contains the dimensions of the smaller resolutionĭigitized_data = np.empty((grid_shape, grid_shape))įor i in range(grid_shape): #row-index of pixel in smaller-resolution gridįor j in range(grid_shape): #column-index of pixel in smaller-resolution grid The digitizing function I've written: import numpy as npįunction_data = 2D array of function values of some 3D shape,Įg.: exp(-(x^2 + y^2 -> want to digitize this The idea is basically the same as changing an HD image to a smaller resolution. I want to split that data into cells, each of which would correspond to a pixel. The photon count data is stored in a 2D array. I want to digitize (= average out over cells) photon count data into pixels given by a grid that tells how they are aligned.
0 Comments
Leave a Reply. |

 RSS Feed
RSS Feed
